Immortalization of human periodontal ligament stem cells by transferring human telomerase reverse transcriptase gene
Introduction
Periodontal disease is one of the main reasons for tooth loss (1). The main goals of current periodontal therapies are to preserve the natural dentition, as well as the esthetics and function of the periodontium (2,3). However, none of the therapies are currently available have shown the ability to completely restore the functionality of all damaged periodontal tissues. Regenerative medicine has certain advantages compared with traditional treatment because it uses tissue engineering and stem cells. Stem cells have high proliferation capacity and multipotent differentiation potential to repair and reconstruct damaged periodontal tissue (4,5).
Human periodontal ligament stem cells (hPDLSCs) were first isolated and named by Seo et al. (6), Who reported that, similar to human dental pulp stem cells (hDPSCs) and bone mesenchymal stem cells (BMSSCs), hPDLSCs also expressed the two mesenchymal stem cell markers STRO-1 and CD146 (7,8). hPDLSCs were also found to have multidirectional differentiation abilities and could be induced into cementoblast-like cells, adipocytes, and collagen-forming cells. When transplanted into immunocompromised mice or rats, hPDLSCs could generate a cementum/PDL-like structure, contributing to periodontal tissue repair (7). Thus, hPDLSCs were considered to be potential seed cells for periodontal tissue engineering.
However, the use of hPDLSCs is often limited, because primary hPDLSCs, in vitro, possess a relatively short replicative lifespan and undergo senescence after several passages (9,10). Therefore, it is necessary to establish a continuous periodontal ligament stem cell line to provide a consistent and homogeneous research cell model.
The hTERT gene lies in the human chromosome 5p15.33 region (11,12). Studies have found that hTERT is a positive regulatory factor in controlling the activity of telomerase, and prevents cell aging through preventing the loss of telomere sequence during mitosis (13-15). Bodnar et al. confirmed that transient hTERT expression in normal human diploid cells could activate telomerase to extend the life of cells (16). Counter et al. also found that transferring the exogenous hTERT gene into human embryonic kidney (HEK) cells could not only immortalize them but the normal function of the cells remains unchanged (17). It is reported that Fujita et al. and Hasegawa et al. had immortalized human periodontal ligament (HPL) cells by transferring the exogenous hTERT gene (18,19), however, they neither separated the hPDLSCs exactly nor detected the carcinogenicity of the HPL cells with exogenous hTERT gene. To our knowledge, a cell line of immortalized hPDLSCs by transferring the exogenous hTERT gene using a lentivirus vector has not been reported yet. Therefore, in this study, the exogenous hTERT gene was transferred into hPDLSCs using a lentivirus vector in an effort to extend cellular lifespan, and hence immortalize this cell line.
Methods
All procedures were carried out in accordance with The Code of Ethics of the World Medical Association (Declaration of Helsinki) for experiments involving human subjects.
hPDLSCs isolation and culture
Premolars extracted for orthodontic reasons were collected from a 12-year-old donor at the Oral Surgery Clinic of the Ninth People’s Hospital affiliated to Shanghai Jiao Tong University School of Medicine (China), with the written informed consent of the patient’s parents. The periodontal ligament was gently separated from the surface of the root and cultured using a modified tissue piece method (7), then the human periodontal ligament cells (hPDLCs) in P2 were isolated by immune magnetic beads separation (beads coated with the antibody of STRO-1) according to the manufacturer's instructions (Miltenyi Biotec, Germany).
Osteogenic and adipogenic differentiation
Osteogenic and adipogenic differentiation was induced as previously reported (20). After differentiation, both types of cells were fixed in 4% paraformaldehyde for 30 min. Cells that underwent osteogenic differentiation were stained with Alizarin red stain for 3–5 min, while cells that underwent adipogenic differentiation were stained with oil red O stain for 10 min. The fixed cells were then observed and images were captured under a light microscope.
Expression of mesenchymal stem cell surface markers
The expression of mesenchymal stem cell surface markers was detected with flow cytometry. hPDLSCs were cultured until they reached 80–90% confluency, and then were detached using a 0.05% trypsin-EDTA solution, washed with PBS and incubated in the dark for 15 min at 4 °C with fluorescence-conjugated monoclonal antibodies for CD73-PE, CD90-FITC, CD105-PE, CD166-PE, CD34-PE, and CD45-FITC (BD Biosciences, Franklin Lakes, NJ, USA). These markers were recommended by the International Society of Cellular Therapy to confirm the mesenchymal phenotype of cells (21). After labeling and washing, cells were fixed in 1% (w/v) paraformaldehyde for 20–30 min, air-dried at 4 °C, sorted using a FACS Calibur flow cytometer and analyzed with FlowJo software (version 7.6.1, Treestar, Ashland, OR, USA). Non-specific fluorescence was measured using specific isotype monoclonal antibodies (BD Biosciences).
Recombinant hTERT lentivirus infection of hPDLSCs
Packaging plasmid PLKO.1 and enveloping plasmid Δ8.9, together with vector pBABE-hygro-hTERT (Addgene, Cambridge, MA, USA) coding for the hTERT gene were used. Production of recombinant hTERT lentivirus was achieved by transfecting HEK293T cells (provided by the Laboratory of the Chinese Academy of Biochemistry and Cell Biology). Supernatant containing viral particles was collected at 24, 48 and 72 h after transfection. For infection of cells, hPDLSCs at 70–80% confluency were exposed to viral supernatant for 24 h, after which the infected cells were selected by culturing in Dulbecco’s modified eagle medium (DMEM, Gibco-BRL, Life Technology, Inc., Grand Island, New York, USA) with hygromycin (25 µg/mL). The selected cells, named hPDLSCs-hTERT, have since been cultured for more than 49 passages. To confirm immortalization and odontoblastic differentiation of the hPDLSCs-hTERT, hPDLSCs-hTERT at passage 49 were used for the following characterization.
To test the stem cells characteristics of hPDLSCs-hTERT (passage 49), the expression of mesenchymal stem cell surface markers was detected with flow cytometry. The markers and protocols were the same as the flow cytometry done with hPDLSCs mentioned above. Furthermore, hPDLSCs-hTERT in passage 49 were cultured in six-well plates at an initial density of 5×105 cells/well in DMEM. After 24 h, the medium was replaced with human mesenchymal stem cell osteogenic/adipogenic differentiation medium (Cyagen Biosciences Inc, Guangzhou, China). At that time, the cells were defined as day 0 (0D). The medium was changed every three days. On day 21 (21D), Alizarin red stain or oil red O was performed to confirm odontoblastic/adipogenic differentiation.
Reverse transcription-polymerase chain reaction (RT-PCR) and real time RT-PCR analysis
To verify whether the hTERT gene was successfully transferred into the hPDLSCs, total RNA isolation, first-strand cDNA synthesis, and PCR processes were performed as reported previously (22); the primer sequences are shown in Table 1. PCR products were separated with agarose gel electrophoresis and observed under UV light.
Table 1
| Gene | Direction | Primer sequences |
|---|---|---|
|
|
Forward | 5'-TTCACCACCATGGAGAAGGCT-3' |
| Reverse | 5'-TCTCATGGTTCACACCCATGA-3' | |
|
|
Forward | 5'-CGGCGACATGGAGAACAAGC-3' |
| Reverse | 5'-AGGTTTTCGCGTGGGTGAGG-3' | |
|
|
Forward | 5'-GCCATTCCAGTTCCTCAAAGC-3' |
| Reverse | 5'-CATGCACCAGGACACCACTT-3' | |
|
|
Forward | 5'-ATGCCTATCACAACAAACC-3' |
| Reverse | 5'-CTCCTTTATGTGACAACTGC-3' | |
|
|
Forward | 5'-GGACCATTCCCACGTCTTCAC-3' |
| Reverse | 5'-CCTTGTAGCCAGGCCCATTG-3' | |
|
|
Forward | 5'-AGGCTGTGTCAGTGAATGCT-3' |
| Reverse | 5'-GTTGCTGAATGTGCATCGGA-3 |
DMP1, dentin matrix protein 1; DSPP, dentin sialophosphoprotein; ALP, alkaline phosphatase; OMD, osteomodulin.
To quantify the expression of the odontoblastic markers dentin sialophosphoprotein (DSPP), dentin matrix protein-1 (DMP1), alkaline phosphatase (ALP), and osteomodulin (OMD) in the hPDLSCs-hTERT (passage 49) before after osteogenic/odontogenic induction, total RNA was isolated from cells using TRIzol Reagent (Invitrogen, Carlsbad, CA, USA). Reverse transcription to cDNA and PCR amplification were then carried out as described previously (23). Primer sequences are shown in Table 1.
Detection of p53 protein by western blotting
hPDLSCs-hTERT or hPDLSCs were harvested at 80–90% confluency, and western blotting was performed after transferring the proteins separated with SDS-PAGE (sodium dodecylsulfate polyacrylamide gel electrophoresis) to Amersham Hybond-ECL membranes (GE Healthcare Bio-Sciences, Piscataway, NJ, USA), as reported previously (24).
Data analysis
All experiments were performed at least in triplicate and reproduced three separate times. Statistical significance was determined using the unpaired student’s t-test. Values of P<0.05 and P<0.01 were accepted as statistically significant.
Results
Isolation and identification of hPDLSCs
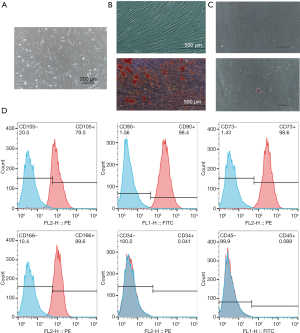
The hPDLSCs were sorted by magnetic-activated cell sorting of generation P2. Cells exhibited a typical fibroblast-like spindle-shaped appearance and had clone-forming ability (Figure 1A). Alizarin red and oil red O staining assays confirmed that the sorted hPDLSCs could be induced to differentiate into osteoblasts and adipocytes (Figure 1B,C). Flow cytometry confirmed that they expressed the mesenchymal stem cell surface markers CD73, CD90, CD105, and CD166, and did not express the endothelial cell marker CD34 or the hematopoietic cell marker CD45 (Figure 1D).
hPDLSCs immortalization
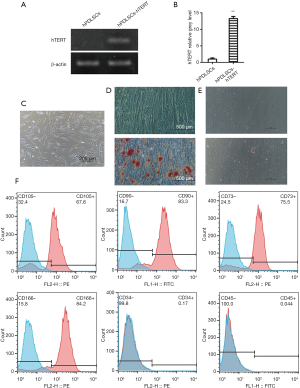
hPDLSCs-hTERT were selected from a 7-day culture in medium with hygromycin. hPDLSCs-hTERT were free of any morphological changes compared with the original hPDLSCs, even at higher passage numbers. The expression of hTERT mRNA was higher in hPDLSCs-hTERT than in hPDLSCs (Figure 2A,B).
Morphologic characteristics of hPDLSCs and hPDLSCs-hTERT.
The morphologic characteristics of both hPDLSCs and hPDLSCs-hTERT appeared stable, with the majority of cells appearing spindle-shaped, with a short spindle and one to two projections, these cells had smaller volumes and central nuclei. The cells adhered to the bottom of the culture dish (Figure 2C), and were able to grow to 80–90% confluency in 3 days when passaged at a 1:4 ratio.
Osteoblastic and adipogenic differentiation of hPDLSCs-hTERT
hPDLSCs-hTERT could be induced to differentiate into osteoblasts within osteogenic differentiation medium, forming mineralized nodules which were positively stained by Alizarin Red (Figure 2D). hPDLSCs-hTERT could also be induced to differentiate into adipocyte within adipogenic differentiation medium, forming lipid droplets were positively stained by oil red O (Figure 2E).
hPDLSCs-hTERT expressed mesenchymal stem cell surface markers
The expression of mesenchymal stem cell surface markers by hPDLSCs-hTERT was analyzed with flow cytometry. Data showed that the same as hPDLSCs, hPDLSCs-hTERT expressed CD73, CD90, CD105, and CD166, but did not express CD34 or CD45 (Figure 2F).
Expression of ALP, DSPP, DMP1, and OMD mRNA in hPDLSCs-hTERT upregulated significantly after odontoblastic/osteogenic induction
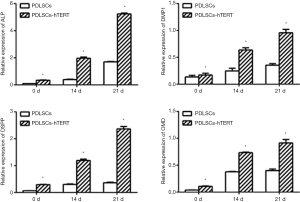
After osteoblastic/odontogenic induction for 14 or 21 days, expression of the odontoblastic markers ALP, DMP1, DSPP, and OMD dramatically increased in both the hPDLSCs and hPDLSCs-hTERT (Figure 3).
p53 expression
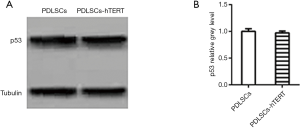
The western blotting results showed that both hPDLSCs-hTERT (passage 49) and hPDLSCs (passage 9) expressed the p53 proteins of 53 KDa (25,26). There was no significant difference between the width and color of the two bands, which demonstrated that hPDLSCs-hTERT had no carcinogenicity (Figure 4A,B).
Discussion
Immortalized cells are able to avoid apoptosis and maintain their characteristic long-term morphology even after numerous passages in culture (18,19). Such cells are thus a good choice for creating a cell model to further study. hPDLSCs could be applied to periodontal regenerative medicine to promote the regeneration of damaged periodontal tissue and to restore periodontal tissue attachments (7). However, due to the aging of hPDLSCs, we need to constantly take the primary hPDLSCs from healthy orthodontic or impacted teeth, which wastes the manpower and resources. Thus, immortalized hPDLSCs could be a promising source of seed cells for periodontal tissue engineering. In this study, the exogenous hTERT gene was transferred into hPDLSCs using a lentivirus vector to extend cellular lifespan, and hence immortalize this cell line, which could make tissue regeneration based on hPDLSCs more convenient.
In this study, the cellular morphology and mesenchymal stem cell surface markers expression of hPDLSCs-hTERT remained similar to the original hPDLSCs. After the addition of human odontoblastic induction medium or human adipogenic induction medium and culture for 21 days, mineralized nodules or lipid droplets could be seen in both hPDLSCs-hTERT (Figure 2D,E) and hPDLSCs (Figure 1B,C). These results indicate that there were no obvious differences between immortalized hPDLSCs-hTERT and hPDLSCs in terms of stem cells characteristics including cell morphology, mesenchymal stem cell surface marker expression, and odontoblasts or adipocytes differentiation ability. This is consistent with previous findings that a cell line immortalized by the exogenous hTERT gene retained many biological characteristics of normal cells, such as cellular morphology, surface features, and biological functions (27,28).
Periodontal membrane is located between alveolar bone and teeth, and the stem cells in the periodontal membrane have the potential to regenerate both them (7). So the main characteristics of hPDLSCs used in tissue engineering are their osteogenic and odontoblastic ability. In most studies, the methods of osteogenic induction and odontogenic induction are the same (18-20,22,29). DSPP is a specific marker for differentiated odontoblasts, which is initially expressed in odontoblasts and hPDLSCs at the bell stage during tooth development (26). Protein encoded by DMP-1 is an ECM protein closely related to osteopontin (OPN) that is expressed by odontoblasts, osteoblasts, and osteocytes, which is critical for dentin formation and periodontal development (29). ALP and OMD are classical markers of osteogenic differentiation (19,20,30). Without odontoblastic/osteogenic induction, hPDLSCs-hTERT expressed the odontogenic-related genes DMP-1, DSPP and osteogenic-related genes ALP, OMD in a relative low level. After 14–21 days induction, the cells expressed high levels of the ALP, DMP-1, OMD and DSPP (Figure 3). These data suggest that the cell line hPDLSCs-hTERT has odontoblastic/osteogenic differentiation potential which making them an ideal seed cell line in periodontal regeneration. Furthermore, we found that the relative expressions of ALP, OMD, DMP-1 and DSPP in hPDLSCs-hTERT are higher than that in hPDLSCs among 0, 14 and 21 days after osteogenic/odontogenic induction (P<0.05). Whether hPDLSCs-hTERT have stronger odontoblastic/osteogenic differentiation potential than hPDLSCs need more experiments to verify.
One of the biggest concerns associated with the use of immortalized cells is tumorigenicity, because tumor cells also have immortal characteristics. The tumor suppressor gene, p53, which lies in the human chromosome 17p13 region, plays a role in slowing or monitoring cell division under normal conditions (26). After p53 gene mutation, cells lose the ability to regulate growth, apoptosis, and DNA repair due to conformational changes (27,28). In the present study, p53 was used to verify whether immortalized PDLSCs were tumorigenic. p53 protein expression showed no significant difference between hPDLSCs-hTERT (passage 49) and hPDLSCs (passage 9) (Figure 4), suggesting that the hPDLSCs-hTERT cell line may have low tumorigenicity.
In conclusion, a line of immortalized hPDLSCs was created in this study by transfecting the exogenous hTERT gene into hPDLSCs, which maintained stem cells’ characteristics. The resulting cell line has osteoblastic/odontoblastic differentiation potential and low tumorigenicity. This cell line could be used as a cell line for studying the periodontal tissue engineering.
Acknowledgments
Funding: This work was supported by the National Natural Science Foundation of China (No.81570964/81371143) and Shanghai Committee of Science and Technology (No. 17140903500).
Footnote
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://fomm.amegroups.org/article/view/10.21037/fomm.2019.09.02/coif). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. All procedures were carried out in accordance with The Code of Ethics of the World Medical Association [Declaration of Helsinki (as revised in 2013)] for experiments involving human subjects. Written informed consent was obtained from the patient’s parents. The institutional ethical approval was waived.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Haas AN, Silva-Boghossian CM, Colombo AP, et al. Predictors of clinical outcomes after periodontal treatment of aggressive periodontitis: 12-month randomized trial. Braz Oral Res 2016; [Crossref] [PubMed]
- Thomas B, Kumari S, Ramitha K, et al. Comparative evaluation of micronutrient status in the serum of diabetes mellitus patients and healthy individuals with periodontitis. J Indian Soc Periodontol 2010;14:46-9. [Crossref] [PubMed]
- Kiss E, Sewon L, Gorzó I, et al. Salivary calcium concentration in relation to periodontal health of female tobacco smokers: a pilot study. Quintessence Int 2010;41:779-85. [PubMed]
- Rattanasuwan K, Lertsukprasert K, Rassameemasmaung S, et al. Long-term outcome following regenerative periodontal treatment of intrabony defects. Odontology 2017;105:191-201. [Crossref] [PubMed]
- Yan X, Xu N, Meng C, et al. Generation of induced pluripotent stem cells from human mesenchymal stem cells of parotid gland origin. Am J Transl Res 2016;8:419-32. [PubMed]
- Seo BM, Miura M, Gronthos S, et al. Investigation of multipotent postnatal stem cells from human periodontal ligament. Lancet 2004;364:149-55. [Crossref] [PubMed]
- Jung IH, Kwon BS, Kim SH, et al. Optimal medium formulation for the long-term expansion and maintenance of human periodontal ligament stem cells. J Periodontol 2013;84:1434-44. [Crossref] [PubMed]
- Zhang QB, Zhang ZQ, Fang SL, et al. Effects of hypoxia on proliferation and osteogenic differentiation of periodontal ligament stem cells: an in vitro and in vivo study. Genet Mol Res 2014;13:10204-14. [Crossref] [PubMed]
- Gamble CM, Barton PA. Baculoviral expression of telomerase in primary human fibroblasts to rejuvenate cells for tissue engineering. J Tissue Eng Regen Med 2012;6:414-20. [Crossref] [PubMed]
- Dabelsteen S, Hercule P, Barron P, et al. Epithelial cells derived from human embryonic stem cells display p16INK4A senescence, hypermotility, and differentiation properties shared by many P63+ somatic cell types. Stem Cells 2009;27:1388-99. [Crossref] [PubMed]
- Bryce LA, Morrisont N, Hoare SF, et al. Mapping of the gene for the human telomerase reverse transcriptase, hTERT, to chromosome 5p15. 33 by fluorescence in situ hybridization. Neoplasia 2000;2:197-201. [Crossref] [PubMed]
- Zhang A, Zheng C, Hou M, et al. Deletion of the telomerase reverse transcriptase gene and haploinsufficiency of telomere maintenance in Cri du chat syndrome. Am J Hum Genet 2003;72:940-8. [Crossref] [PubMed]
- Zhang H, Huang Y, Wang L, et al. Immortalization of porcine placental trophoblast cells through reconstitution of telomerase activity. Theriogenology 2016;85:1446-56. [Crossref] [PubMed]
- Hills M, Lansdorp PM. Short telomeres resulting from heritable mutations in the telomerase reverse transcriptase gene predispose for a variety of malignancies. Ann N Y Acad Sci 2009;1176:178-90. [Crossref] [PubMed]
- Zhang RG, Zhao JJ, Yang LQ, et al. RNA interference-mediated hTERT inhibition enhances TRAIL-induced apoptosis in resistant hepatocellular carcinoma cells. Oncol Rep 2010;23:1013-9. [PubMed]
- Bodnar AG, Ouellette M, Frolkis M, et al. Extension of life-span by introduction of telomerase into normal human cells. Science 1998;279:349-52. [Crossref] [PubMed]
- Counter CM, Hahn WC, Wei W, et al. Dissociation among in vitro telomerase activity, telomere maintenance, and cellular immortalization. Proc Natl Acad Sci U S A 1998;95:14723-8. [Crossref] [PubMed]
- Fujita T, Otsuka-Tanaka Y, Tahara H, et al. Establishment of immortalized clonal cells derived from periodontal ligament cells by induction of the hTERT gene. J Oral Sci 2005;47:177-84. [Crossref] [PubMed]
- Hasegawa T, Chosa N, Asakawa T, et al. Establishment of immortalized human periodontal ligament cells derived from deciduous teeth. Int J Mol Med 2010;26:701-5. [Crossref] [PubMed]
- Chadipiralla K, Yochim J M, Bahuleyan B, et al. Osteogenic differentiation of stem cells derived from human periodontal ligaments and pulp of human exfoliated deciduous teeth. Cell Tissue Res 2010;340:323-33. [Crossref] [PubMed]
- Dominici M, Le Blanc K, Mueller I, et al. Minimal criteria for defining multipotent mesenchymal stromal cells. The International Society for Cellular Therapy position statement. Cytotherapy 2006;8:315-7. [Crossref] [PubMed]
- Gronthos S, Brahim J, Li W, et al. Stem cell properties of human dental pulp stem cells. J Dent Res 2002;81:531-5. [Crossref] [PubMed]
- Bustin SA, Nolan T. Pitfalls of quantitative real-time reverse-transcription polymerase chain reaction. J Biomol Tech 2004;15:155-66. [PubMed]
- de Almeida VH, de Melo AC, Meira DD, et al. Radiotherapy modulates expression of EGFR, ERCC1 and p53 in cervical cancer. Braz J Med Biol Res 2017;51:e6822 [Crossref] [PubMed]
- Yu J, Liu X, Ye H, et al. Genomic characterization of the human mitochondrial tumor suppressor gene 1 (MTUS1): 5'cloning and preliminary analysis of the multiple gene promoters. BMC Res Notes 2009;2:109. [Crossref] [PubMed]
- Amariglio N, Hirshberg A, Scheithauer B W, et al. Donor-derived brain tumor following neural stem cell transplantation in an ataxia telangiectasia patient. PLoS Med 2009;6:e1000029 [Crossref] [PubMed]
- Wege H, Heim D, Lütgehetmann M, et al. Forced activation of β-catenin signaling supports the transformation of hTERT-immortalized human fetal hepatocytes. Mol Cancer Res 2011;9:1222-31. [Crossref] [PubMed]
- Huang G, Zheng Q, Sun J, et al. Stabilization of cellular properties and differentiation mutilpotentialof human mesenchymal stem cells transduced with hTERT gene in a long-term culture. J Cell Biochem 2008;103:1256-69. [Crossref] [PubMed]
- Nowwarote N, Sukarawan W, Pavasant P, et al. Basic fibroblast growth factor regulates phosphate/pyrophosphate regulatory genes in stem cells isolated from human exfoliated deciduous teeth. Stem Cell Res Ther 2018;9:345. [Crossref] [PubMed]
- Ulrich C, Rolauffs B, Abele H, et al. Low osteogenic differentiation potential of placenta-derived mesenchymal stromal cells correlates with low expression of the transcription factors Runx2 and Twist2. Stem Cells Dev 2013;22:2859-72. [Crossref] [PubMed]
Cite this article as: Gao L, Zhao F, Jiang W, Niu C, Yuan K, Lin W, Hu X, Ma R, Huang Z. Immortalization of human periodontal ligament stem cells by transferring human telomerase reverse transcriptase gene. Front Oral Maxillofac Med 2019;1:6.